6 results
Impact of measurement and feedback on chlorhexidine gluconate bathing among intensive care unit patients: A multicenter study
- Yoona Rhee, Mary K. Hayden, Michael Schoeny, Arthur W. Baker, Meghan A. Baker, Shruti Gohil, Chanu Rhee, Naasha J. Talati, David K. Warren, Sharon Welbel, Karen Lolans, Bardia Bahadori, Pamela B. Bell, Heilen Bravo, Thelma Dangana, Christine Fukuda, Tracey Habrock Bach, Alicia Nelson, Andrew T. Simms, Pam Tolomeo, Robert Wolf, Rachel Yelin, Michael Y. Lin, for the CDC Prevention Epicenters Program
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 44 / Issue 9 / September 2023
- Published online by Cambridge University Press:
- 13 September 2023, pp. 1375-1380
- Print publication:
- September 2023
-
- Article
- Export citation
-
Objective:
To assess whether measurement and feedback of chlorhexidine gluconate (CHG) skin concentrations can improve CHG bathing practice across multiple intensive care units (ICUs).
Design:A before-and-after quality improvement study measuring patient CHG skin concentrations during 6 point-prevalence surveys (3 surveys each during baseline and intervention periods).
Setting:The study was conducted across 7 geographically diverse ICUs with routine CHG bathing.
Participants:Adult patients in the medical ICU.
Methods:CHG skin concentrations were measured at the neck, axilla, and inguinal region using a semiquantitative colorimetric assay. Aggregate unit-level CHG skin concentration measurements from the baseline period and each intervention period survey were reported back to ICU leadership, which then used routine education and quality improvement activities to improve CHG bathing practice. We used multilevel linear models to assess the impact of intervention on CHG skin concentrations.
Results:We enrolled 681 (93%) of 736 eligible patients; 92% received a CHG bath prior to survey. At baseline, CHG skin concentrations were lowest on the neck, compared to axillary or inguinal regions (P < .001). CHG was not detected on 33% of necks, 19% of axillae, and 18% of inguinal regions (P < .001 for differences in body sites). During the intervention period, ICUs that used CHG-impregnated cloths had a 3-fold increase in patient CHG skin concentrations as compared to baseline (P < .001).
Conclusions:Routine CHG bathing performance in the ICU varied across multiple hospitals. Measurement and feedback of CHG skin concentrations can be an important tool to improve CHG bathing practice.
Multicenter evaluation of contamination of the healthcare environment near patients with Candida auris skin colonization – ERRATUM
- Sarah E. Sansom, Gabrielle M. Gussin, Raveena D Singh, Pamela B Bell, Ellen Benson, Jinal Makhija, Mary Carl Froilan, Raheeb Saavedra, Robert Pedroza, Christine Thotapalli, Christine Fukuda, Ellen Gough, Stefania Marron, Maria Del Mar Villanueva Guzman, Julie A. Shimabukuro, Lydia Mikhail, Stephanie Black, Massimo Pacilli, Hira Adil, Cassiana E. Bittencourt, Matthew Zahn, Nicholas Moore, D. Sexton, Judith Noble-Wang, Meghan Lyman, Michael Lin, Susan Huang, Mary K. Hayden
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 2 / Issue 1 / 2022
- Published online by Cambridge University Press:
- 07 October 2022, e166
-
- Article
-
- You have access Access
- Open access
- HTML
- Export citation
Multicenter evaluation of contamination of the healthcare environment near patients with Candida auris skin colonization
- Sarah Sansom, Gabrielle M. Gussin, Raveena D Singh, Pamela B Bell, Ellen Benson Jinal, Makhija, Raheeb Froilan, Raheeb Saavedra, Robert Pedroza, Christine Thotapalli, Christine Fukuda, Ellen Gough, Stefania Marron, Maria Del Mar Villanueva Guzman, Julie A. Shimabukuro, Lydia Mikhail, Stephanie Black, Massimo Pacilli, Hira Adil, Cassiana E. Bittencourt, Matthew Zahn, Nicholas Moore, D. Joseph Sexton, Judith Noble-Wang, Meghan Lyman, Michael Lin, Susan Huang, Mary Hayden
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 2 / Issue S1 / July 2022
- Published online by Cambridge University Press:
- 16 May 2022, pp. s78-s79
-
- Article
-
- You have access Access
- Open access
- Export citation
-
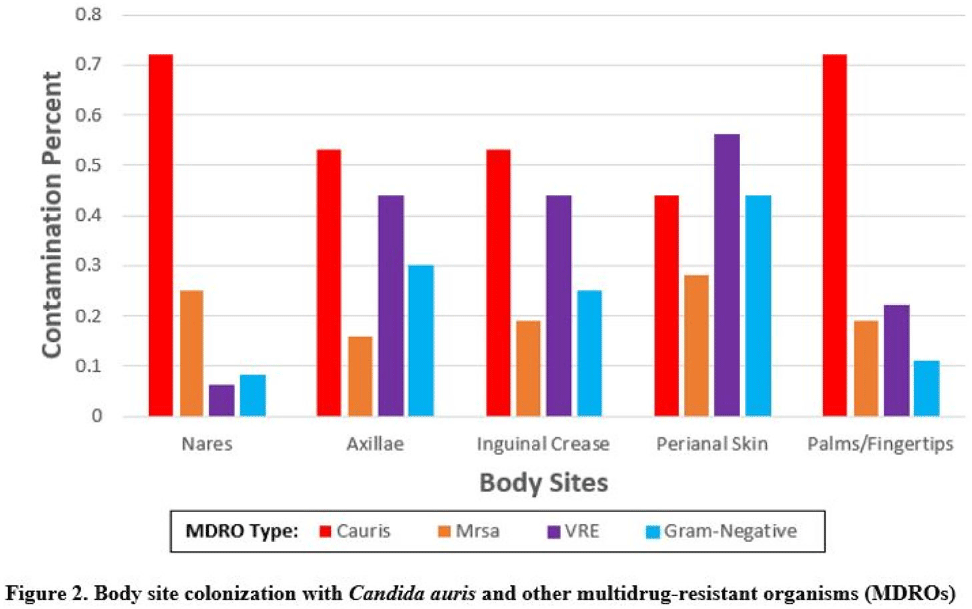
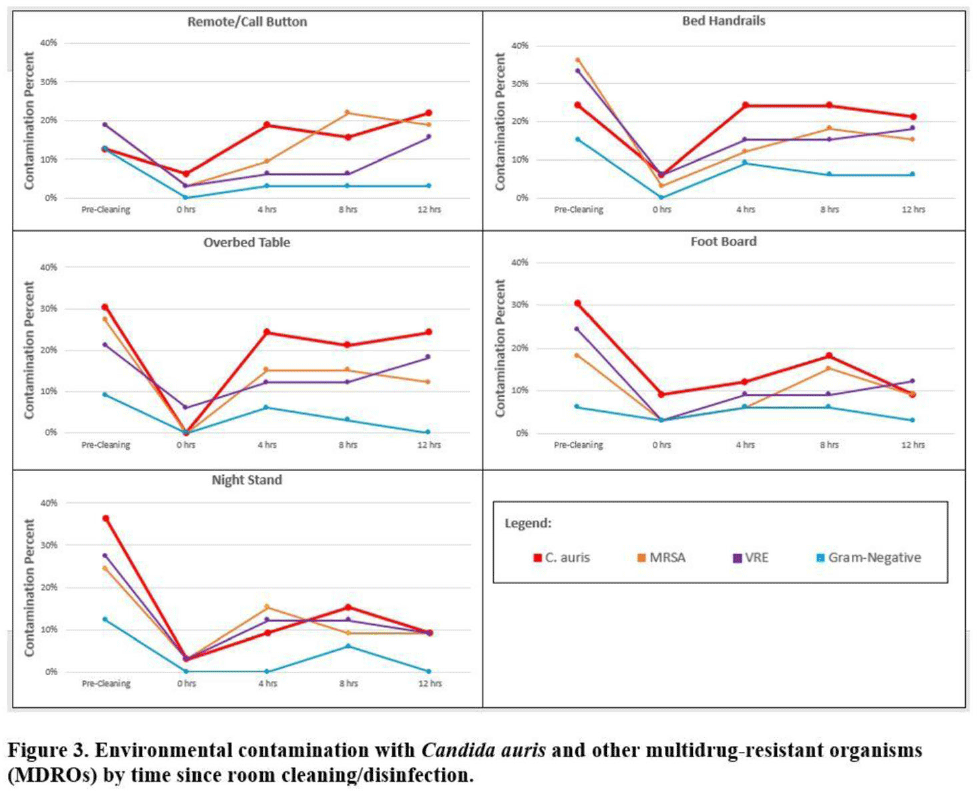
Background:Candida auris is an emerging multidrug-resistant yeast that is transmitted in healthcare facilities and is associated with substantial morbidity and mortality. Environmental contamination is suspected to play an important role in transmission but additional information is needed to inform environmental cleaning recommendations to prevent spread. Methods: We conducted a multiregional (Chicago, IL; Irvine, CA) prospective study of environmental contamination associated with C. auris colonization of patients and residents of 4 long-term care facilities and 1 acute-care hospital. Participants were identified by screening or clinical cultures. Samples were collected from participants’ body sites (eg, nares, axillae, inguinal creases, palms and fingertips, and perianal skin) and their environment before room cleaning. Daily room cleaning and disinfection by facility environmental service workers was followed by targeted cleaning of high-touch surfaces by research staff using hydrogen peroxide wipes (see EPA-approved product for C. auris, List P). Samples were collected immediately after cleaning from high-touch surfaces and repeated at 4-hour intervals up to 12 hours. A pilot phase (n = 12 patients) was conducted to identify the value of testing specific high-touch surfaces to assess environmental contamination. High-yield surfaces were included in the full evaluation phase (n = 20 patients) (Fig. 1). Samples were submitted for semiquantitative culture of C. auris and other multidrug-resistant organisms (MDROs) including methicillin-resistant Staphylococcus aureus (MRSA), vancomycin-resistant Enterococcus (VRE), extended-spectrum β-lactamase–producing Enterobacterales (ESBLs), and carbapenem-resistant Enterobacterales (CRE). Times to room surface contamination with C. auris and other MDROs after effective cleaning were analyzed. Results:Candida auris colonization was most frequently detected in the nares (72%) and palms and fingertips (72%). Cocolonization of body sites with other MDROs was common (Fig. 2). Surfaces located close to the patient were commonly recontaminated with C. auris by 4 hours after cleaning, including the overbed table (24%), bed handrail (24%), and TV remote or call button (19%). Environmental cocontamination was more common with resistant gram-positive organisms (MRSA and, VRE) than resistant gram-negative organisms (Fig. 3). C. auris was rarely detected on surfaces located outside a patient’s room (1 of 120 swabs; <1%). Conclusions: Environmental surfaces near C. auris–colonized patients were rapidly recontaminated after cleaning and disinfection. Cocolonization of skin and environment with other MDROs was common, with resistant gram-positive organisms predominating over gram-negative organisms on environmental surfaces. Limitations include lack of organism sequencing or typing to confirm environmental contamination was from the room resident. Rapid recontamination of environmental surfaces after manual cleaning and disinfection suggests that alternate mitigation strategies should be evaluated.
Funding: None
Disclosures: None
Indwelling medical devices and skin microorganisms on ICU patients bathed with chlorhexidine gluconate
- Yoona Rhee, Mary Hayden, Michael Schoeny, Christine Fukuda, Pamela B. Bell, Andrew Simms, Beverly Sha, Carlos Santos, Michael Lin
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 2 / Issue S1 / July 2022
- Published online by Cambridge University Press:
- 16 May 2022, pp. s43-s44
-
- Article
-
- You have access Access
- Open access
- Export citation
-
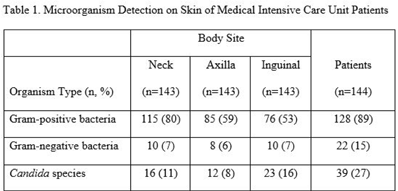
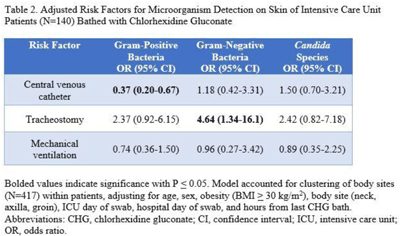
Background: Bathing ICU patients with chlorhexidine gluconate (CHG) decreases bloodstream infections and multidrug-resistant organism transmission. The efficacy of CHG bathing on skin microorganism reduction may be influenced by patient-level clinical factors. We assessed the impact of clinical factors on the recovery of microorganisms from the skin of patients admitted to an ICU who were receiving routine CHG bathing. Methods: We analyzed data obtained from 6 single-day point-prevalence surveys of adult ICU patients between January and October 2018 at 1 medical ICU, in the context of a CHG bathing quality initiative. Demographics and covariates were collected at the bedside and by chart review. Skin swabs were collected from neck, axilla, and inguinal regions and were plated to selective and nonselective media. Standard microbiologic methods were used for species identification and susceptibilities. Multivariable models included patients who received a CHG bath and accounted for clustering of body sites within patients. Results: Across all time points, 144 patients participated, yielding 429 skin swab samples. Mean age was 57 years (SD, 17); 49% were male; 44% had a central venous catheter; and 15% had a tracheostomy Also, 140 patients (97%) had >1 CHG bath prior to skin swab collection, with a median of 9 hours since their last CHG bath (IQR, 6–13 hours). Gram-positive bacteria were more commonly recovered than gram-negative or Candida spp across all skin sites (Table 1). Variation by body site was detected only for gram-positive bacteria, with recovery more common from the neck compared to axilla or groin sites. On multivariate logistic regression (Table 2), presence of central venous catheter was associated with lower odds of gram-positive bacteria recovery among those who received a CHG bath. Presence of tracheostomy was associated with a significantly higher odds of gram-negative bacteria detection on skin. No clinical factors were independently associated with recovery of Candida spp. Conclusions: Central venous catheter presence was associated with lower odds of gram-positive bacteria detection on skin, suggesting the possibility of higher quality CHG bathing among such patients. Tracheostomy presence was associated with greater odds of gram-negative bacteria detection, suggesting that it may be a potential reservoir for skin contamination or colonization. Indwelling medical devices may influence CHG bathing effectiveness in reducing microorganism burden on skin.
Funding: None
Disclosures: None
Genomic investigation to identify the source of SARS-CoV-2 infection among healthcare personnel
- Sarah Sansom, Hannah Barbian, Evan Snitkin, Christine Fukuda, Nicholas Moore, Lahari Thotapalli, Elias Baied, DO Young Kim, Mary Hayden, Michael Lin
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 2 / Issue S1 / July 2022
- Published online by Cambridge University Press:
- 16 May 2022, pp. s74-s75
-
- Article
-
- You have access Access
- Open access
- Export citation
-
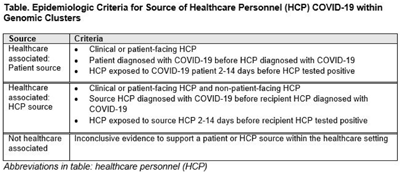
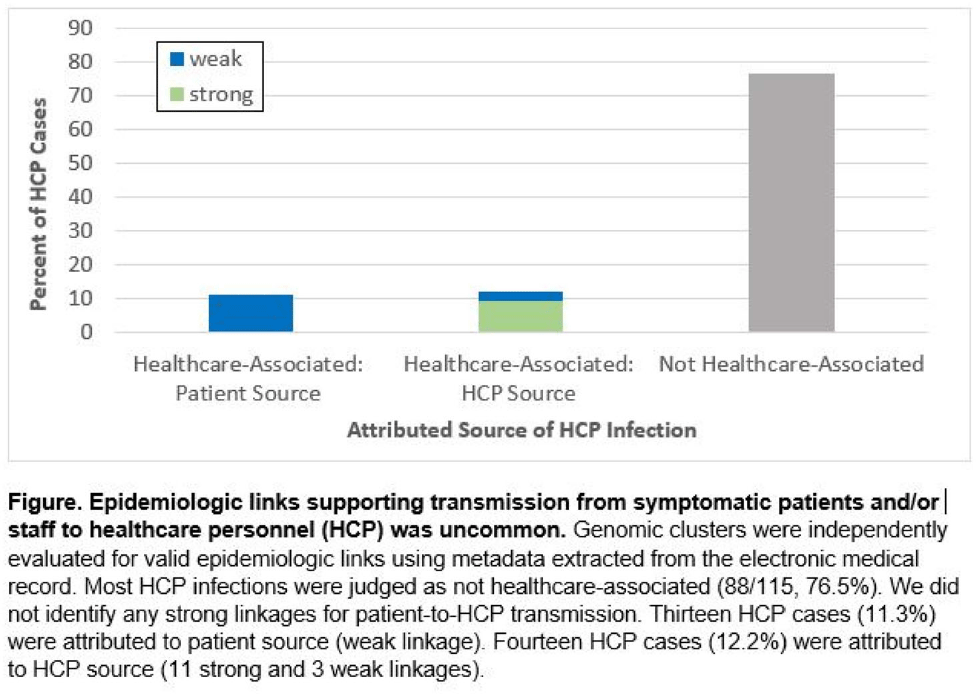
Background: Contact tracing alone is often inadequate to determine the source of healthcare personnel (HCP) COVID-19 when SARS-CoV-2 is widespread in the community. We combined whole-genome sequencing (WGS) with traditional epidemiologic analysis to investigate the frequency with which patients or other HCP with symptomatic COVID-19 acted as the source of HCP infection at a large tertiary-care center early in the pandemic. Methods: Cohort samples were selected from patients and HCP with PCR-positive SARS-CoV-2 infection from a period with complete retention of samples (March 14, 2021–April 10, 2020) at Rush University Medical Center, a 664-bed hospital in Chicago, Illinois. During this period, testing was limited to symptomatic patients and HCP. Recommended respiratory equipment for HCP evolved under guidance, including a 19-day period when medical face masks were recommended for COVID-19 care except for aerosol-generating procedures. Viral RNA was extracted and sequenced (NovaSeq, Illumina) from remnant nasopharyngeal swab samples in M4RT viral transport medium. Genomes with >90% coverage underwent cluster detection using a 2 single-nucleotide variant genetic distance cutoff. Genomic clusters were independently evaluated for valid epidemiologic links by 2 infectious diseases physicians (with a third adjudicator) using metadata extracted from the electronic medical record and according to predetermined criteria (Table 1). Results: In total, 1,031 SARS-CoV-2 sequences were analyzed, identifying 49 genomic clusters with HCP (median, 8; range, 2–43 members per cluster; total, 268 patients and 115 HCP) (Fig. 1). Also, 20,190 flowsheet activities were documented for cohort HCP and patient interactions, including 686 instances in which a cohort HCP contributed to a cohort patient’s chart. Most HCP infections were considered not healthcare associated (88 of 115, 76.5%). We did not identify any strong linkages for patient-to-HCP transmission. Moreover, 13 HCP cases (11.3%) were attributed to patient source (weak linkage). Also, 14 HCP cases (12.2%) were attributed to HCP source (11 strong and 3 weak linkages). Weak linkages were due to lack of epidemiologic data for HCP location, particularly nonclinical staff (eg, an environmental service worker who lacked location documentation to rule out patient-specific contact). Agreement for epidemiologic linkage between the 2 evaluators was high (κ, 0.91). Conclusions: Using genomic and epidemiologic data, we found that most HCP COVID-19 infections were not healthcare associated. We found weak evidence to support symptomatic patient-to-HCP transmission of SARS-CoV-2 and stronger evidence for HCP-to-HCP transmission. Large genomic clusters without plausible epidemiologic links were identified, reflecting the limited utility of genomic surveillance alone to characterize chains of transmission of SARS-CoV-2 during extensive community spread.
Funding: None
Disclosures: None

Admission and Discharge Sampling Underestimates Multidrug-Resistant Organism (MDRO) Acquisition in an Intensive Care Unit
- Sarah Sansom, Michael Lin, Christine Fukuda, Teppei Shimasaki, Thelma Dangana, Nicholas Moore, Rachel Yelin, Yoona Rhee, Lina Tabith, Jianrong Sheng, Enrique Cornejo Cisneros, John Murray, Kyle Chang, Karen Lolans, Michelle Ariston, William Rotunno, Hazel Ramos, Haiying Li, Khaled Aboushaala, Naomi Iwai, Christine Bassis, Vincent Young, Mary Hayden
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 1 / Issue S1 / July 2021
- Published online by Cambridge University Press:
- 29 July 2021, p. s28
-
- Article
-
- You have access Access
- Open access
- Export citation
-
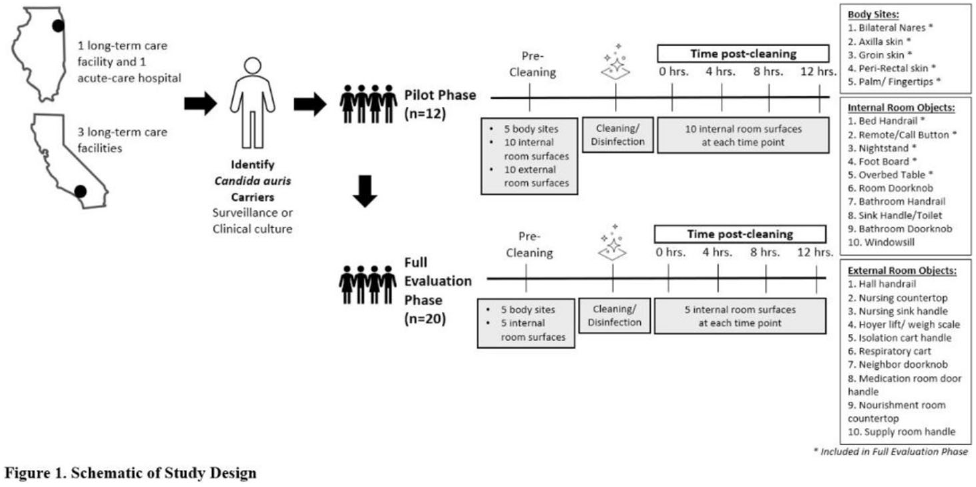
Background: Identification of hospitalized patients with enteric multidrug-resistant organism (MDRO) carriage, combined with implementation of targeted infection control interventions, may help reduce MDRO transmission. However, the optimal surveillance approach has not been defined. We sought to determine whether daily serial rectal surveillance for MDROs detects more incident cases (acquisition) of MDRO colonization in medical intensive care unit (MICU) patients than admission and discharge surveillance alone. Methods: Prospective longitudinal observational single-center study from January 11, 2017, to January 11, 2018. Inclusion criteria were ≥3 consecutive MICU days and ≥2 rectal or stool swabs per MICU admission. Daily rectal or stool swabs were collected from patients and cultured for MDROs, including vancomycin-resistant Enterococcus (VRE), carbapenem-resistant Enterobacterales (CRE), third-generation cephalosporin-resistant Enterobacterales (3GCR), and extended-spectrum β-lactamase–producing Enterobacterales (ESBL-E) (as a subset of 3GCR). MDRO detection at any time during the MICU stay was used to calculate prevalent colonization. Incident colonization (acquisition) was defined as new detection of an MDRO after at least 1 prior negative swab. We then determined the proportion of prevalent and incident cases detected by daily testing that were also detected when only first swabs (admission) and last swabs (discharge) were tested. Data were analyzed using SAS version 9.4 software. Results: In total, 939 MICU stays of 842 patients were analyzed. Patient characteristics were median age 64 years (interquartile range [IQR], 51–74), median MICU length of stay 5 days (IQR, 3–8), median number of samples per admission 3 (IQR, 2–5), and median Charlson index 4 (IQR, 2–7). Prevalent colonization with any MDRO was detected by daily swabbing in 401 stays (42.7%). Compared to daily serial swabbing, an admission- and discharge-only approach detected ≥86% of MDRO cases (ie, overall prevalent MDRO colonization). Detection of incident MDRO colonization by an admission- or discharge-only approach would have detected fewer cases than daily swabbing (Figure 1); ≥34% of total MDRO acquisitions would have been missed. Conclusions: Testing patients upon admission and discharge to an MICU may fail to detect MDRO acquisition in more than one-third of patients, thereby reducing the effectiveness of MDRO control programs that are targeted against known MDRO carriers. The poor performance of a single discharge swab may be due to intermittent or low-level MDRO shedding, inadequate sampling, or transient MDRO colonization. Additional research is needed to determine the optimal surveillance approach of enteric MDRO carriage.
Funding: No
Disclosures: None
Figure 1.